Developer Guide¶
QUICK API¶
Starting from version 20.06, the QUICK build system compiles the source code and creates static or shared object libraries. Such libraries are then linked to the main QUICK program. Assuming that a user specifies a prefix ($installdir) during the configuration of legacy or CMake builds, libraries will be located inside $installdir/lib.
In the case of the legacy build system, there will be subdirectories $installdir/lib/$buildtype where buildtype could be serial, mpi or cuda. Required .mod or header files can be found inside $QUICK_HOME/include/$buildtype.
In the case of the CMake build system the libraries will be in directory $installdir/lib. Depending on the type of build, there will be the serial library libquick.so and any of CUDA and MPI enabled libraries libquick_cuda.so, libquick_mpi.so, and libquick_mpi_cuda.so. The corresponding module files are located in build type specific subdirectories of $installdir/include/libxc and $installdir/include/quick.
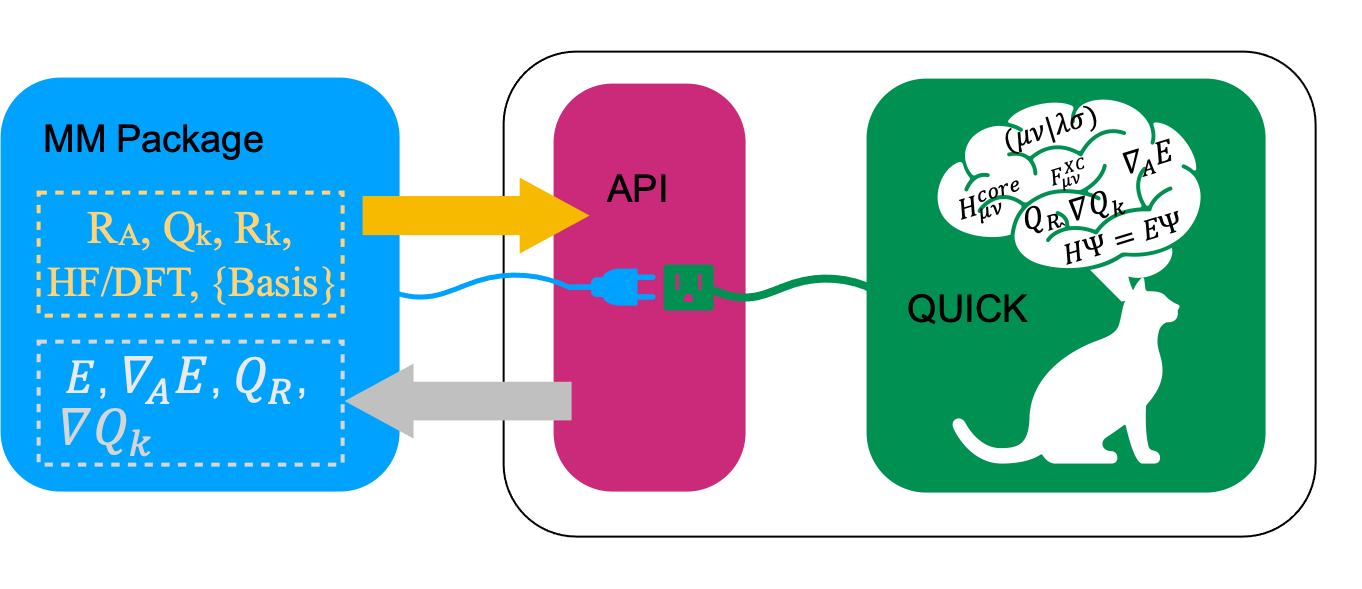
It is possible to link QUICK libraries into external programs to obtain HF/DFT energies, gradients and point charge gradients through the Fortran 90 QUICK API. This is useful for example for MM programs to perform QM/MM calculations. We will explain the usage of the API with an example.
Let us consider a simple system containing a water molecule surrounded by 3 point charges. We now create the following fortran module (test_module.f90) and store atomic coordinates and charges for 5 snapshots. Furthermore, we implement several subroutines to load test data and print data retrieved from QUICK.
! Test module for QUICK API
module test_quick_api_module
implicit none
private
public :: loadTestData, printQuickOutput
#ifdef MPIV
public :: mpi_initialize, printQuickMPIOutput, mpi_exit
#endif
! A test system with one water molecule and 3 point charges.
! Atomic coordinates, external point charges and their coordinates
! for five snapshots.
double precision, dimension(1:45) :: all_coords
double precision, dimension(1:60) :: all_extchg
data all_coords &
/-0.778803, 0.000000, 1.132683, &
-0.666682, 0.764099, 1.706291, &
-0.666682,-0.764099, 1.706290, &
-0.678803, 0.000008, 1.232683, &
-0.724864, 0.755998, 1.606291, &
-0.724862,-0.756005, 1.606290, &
-0.714430, 0.000003, 1.267497, &
-0.687724, 0.761169, 1.624424, &
-0.687723,-0.761172, 1.624427, &
-0.771504, 0.000000, 1.167497, &
-0.669068, 0.763767, 1.697008, &
-0.669068,-0.763767, 1.697008, &
-0.771372, 0.000000, 1.162784, &
-0.668845, 0.767538, 1.698983, &
-0.668845,-0.767538, 1.698982/
data all_extchg &
/1.6492, 0.0000,-2.3560, -0.8340, &
0.5448, 0.0000,-3.8000, 0.4170, &
0.5448, 0.0000,-0.9121, 0.4170, &
1.6492, 0.0000,-2.3560, -0.8360, &
0.5448, 0.0000,-3.8000, 0.4160, &
0.5448, 0.0000,-0.9121, 0.4160, &
1.6492, 0.0000,-2.3560, -0.8380, &
0.5448, 0.0000,-3.8000, 0.4150, &
0.5448, 0.0000,-0.9121, 0.4150, &
1.6492, 0.0000,-2.3560, -0.8400, &
0.5448, 0.0000,-3.8000, 0.4140, &
0.5448, 0.0000,-0.9121, 0.4140, &
1.6492, 0.0000,-2.3560, -0.8420, &
0.5448, 0.0000,-3.8000, 0.4130, &
0.5448, 0.0000,-0.9121, 0.4130/
! number of point charges per frame
integer :: nptg_pframe = 3
interface loadTestData
module procedure load_test_data
end interface loadTestData
contains
subroutine load_test_data(frame, natoms, nxt_charges, coord, xc_coord)
implicit none
integer, intent(in) :: frame, natoms, nxt_charges
double precision, intent(inout) :: coord(3, natoms)
double precision, intent(out) :: xc_coord(4, nxt_charges)
integer :: i, j, k
k=natoms*3*(frame-1) + 1
do i=1,natoms
do j=1,3
coord(j,i) = all_coords(k)
k=k+1
enddo
enddo
if(nxt_charges>0) then
k=nptg_pframe*4*(frame-1) + 1
do i=1,nxt_charges
do j=1,4
xc_coord(j,i) = all_extchg(k)
k=k+1
enddo
enddo
endif
end subroutine load_test_data
#ifdef MPIV
! Initialize mpi library and save mpirank and mpisize.
subroutine mpi_initialize(mpisize, mpirank, master, mpierror)
implicit none
integer, intent(inout) :: mpisize, mpirank, mpierror
logical, intent(inout) :: master
include 'mpif.h'
call MPI_INIT(mpierror)
call MPI_COMM_RANK(MPI_COMM_WORLD,mpirank,mpierror)
call MPI_COMM_SIZE(MPI_COMM_WORLD,mpisize,mpierror)
call MPI_BARRIER(MPI_COMM_WORLD,mpierror)
if(mpirank .eq. 0) then
master = .true.
else
master = .false.
endif
end subroutine mpi_initialize
! Prints mpi output sequentially.
subroutine printQuickMPIOutput(natoms, nxt_charges, atomic_numbers, &
totEne, gradients, ptchg_grad, mpirank)
implicit none
integer, intent(in) :: natoms, nxt_charges, mpirank
integer, intent(in) :: atomic_numbers(natoms)
double precision, intent(in) :: totEne
double precision, intent(in) :: gradients(3,natoms)
double precision, intent(in) :: ptchg_grad(3,nxt_charges)
write(*,*) ""
write(*,'(A11, 1X, I3, 1x, A3)') "--- MPIRANK", mpirank, "---"
write(*,*) ""
call printQuickOutput(natoms, nxt_charges, atomic_numbers, totEne, &
gradients, ptchg_grad)
end subroutine printQuickMPIOutput
subroutine mpi_exit
implicit none
integer :: mpierror
include 'mpif.h'
call MPI_FINALIZE(mpierror)
call exit(0)
end subroutine mpi_exit
#endif
subroutine printQuickOutput(natoms, nxt_charges, atomic_numbers, totEne, &
gradients, ptchg_grad)
implicit none
integer, intent(in) :: natoms, nxt_charges
integer, intent(in) :: atomic_numbers(natoms)
double precision, intent(in) :: totEne
double precision, intent(in) :: gradients(3,natoms)
double precision, intent(in) :: ptchg_grad(3,nxt_charges)
integer :: i, j
! Print energy
write(*,*) ""
write(*,*) "*** TESTING QUICK API ***"
write(*,*) ""
write(*,*) "PRINTING ENERGY"
write(*,*) "---------------"
write(*,*) ""
write(*, '(A14, 3x, F14.10, 1x, A4)') "TOTAL ENERGY =",totEne,"A.U."
! Print gradients
write(*,*) ""
write(*,*) "PRINTING GRADIENTS"
write(*,*) "------------------"
write(*,*) ""
write(*, '(A14, 3x, A6, 10x, A6, 10x, A6)') "ATOMIC NUMBER","GRAD-X","GRAD-Y","GRAD-Z"
do i=1,natoms
write(*,'(6x, I5, 2x, F14.10, 2x, F14.10, 2x, F14.10)') atomic_numbers(i), &
gradients(1,i), gradients(2,i), gradients(3,i)
enddo
! Print point charge gradients
if(nxt_charges>0) then
write(*,*) ""
write(*,*) "PRINTING POINT CHARGE GRADIENTS"
write(*,*) "-------------------------------"
write(*,*) ""
write(*, '(A14, 3x, A6, 10x, A6, 10x, A6)') "CHARGE NUMBER","GRAD-X","GRAD-Y","GRAD-Z"
do i=1,nxt_charges
write(*,'(6x, I5, 2x, F14.10, 2x, F14.10, 2x, F14.10)') i, ptchg_grad(1,i), &
ptchg_grad(2,i), ptchg_grad(3,i)
enddo
endif
write(*,*) ""
end subroutine printQuickOutput
end module
Next, we implement the following example program (example.f90) that uses the above module and calls QUICK through the API.
! Program for testing QUICK API
program test_quick_api
use test_quick_api_module, only : loadTestData, printQuickOutput
use quick_api_module, only : setQuickJob, getQuickEnergy, &
getQuickEnergyGradients, deleteQuickJob
use quick_exception_module
#ifdef MPIV
use test_quick_api_module, only : mpi_initialize, printQuickMPIOutput, mpi_exit
use quick_api_module, only : setQuickMPI
#endif
implicit none
#ifdef MPIV
include 'mpif.h'
#endif
! i, j are some integers useful for loops, frames is the number of
! test snapshots (md steps), ierr is for error handling
integer :: i, j, frames, ierr
! number of atoms, number of atom types, number of external point charges
integer :: natoms, nxt_charges
! atom type ids, atomic numbers, atomic coordinates, point charges and
! coordinates
integer, allocatable, dimension(:) :: atomic_numbers
double precision, allocatable, dimension(:,:) :: coord
double precision, allocatable, dimension(:,:) :: xc_coord
! name of the quick template input file
character(len=80) :: fname
! job card
character(len=200) :: keywd
! total qm energy, mulliken charges, gradients and point charge gradients
double precision :: totEne
double precision, allocatable, dimension(:,:) :: gradients
double precision, allocatable, dimension(:,:) :: ptchgGrad
#ifdef MPIV
! essential mpi information
integer :: mpierror = 0
integer :: mpirank = 0
integer :: mpisize = 1
logical :: master = .true.
! Initialize mpi library and get mpirank, mpisize
call mpi_initialize(mpisize, mpirank, master, mpierror)
! Setup quick mpi using api, called only once
call setQuickMPI(mpirank,mpisize,ierr)
#endif
! Set molecule size. We consider a water molecule surounded by 3 point
! charges in this test case.
natoms = 3
nxt_charges = 3
! We consider 5 snapshots of this test system (mimics 5 md steps).
frames = 5
! Alocate memory for some input and output arrays.
if ( .not. allocated(atomic_numbers)) allocate(atomic_numbers(natoms), stat=ierr)
if ( .not. allocated(coord)) allocate(coord(3,natoms), stat=ierr)
if ( .not. allocated(gradients)) allocate(gradients(3,natoms), stat=ierr)
! Fill up memory with test values, coordinates and external charges will be loded inside
! the loop below.
fname = 'api_water_rhf_631g'
keywd = 'HF BASIS=6-31G CUTOFF=1.0D-10 DENSERMS=1.0D-6 GRADIENT EXTCHARGES'
!keywd =''
atomic_numbers(1) = 8
atomic_numbers(2) = 1
atomic_numbers(3) = 1
! Set the gradient vector to zero.
gradients = 0.0d0
! initialize QUICK, required only once. Assumes keywords for
! the QUICK job are provided through a template file.
call setQuickJob(fname, keywd, natoms, atomic_numbers, ierr)
do i=1, frames
! Actual QM/MM simulations may have different number of point charges during MD.
! Use this trick to mimic this & load coordinates and external point charges for ith step.
nxt_charges = mod(i,4)
! Allocate memory for xyz coordinates of the point charges and gradients.
! Note that in xc_coord array, the first 3 columns are the xyz coordinates
! of the point charges and fourth column is the charge.
if ( .not. allocated(xc_coord)) allocate(xc_coord(4,nxt_charges), stat=ierr)
if ( .not. allocated(ptchgGrad)) allocate(ptchgGrad(3,nxt_charges), stat=ierr)
! Set the point charge gradient vector to zero.
ptchgGrad = 0.0d0
! Load test data.
call loadTestData(i, natoms, nxt_charges, coord, xc_coord)
! Compute required quantities, call only a or b.
! a. compute energy
! call getQuickEnergy(coord, nxt_charges, xc_coord, totEne)
! b. Compute energies, gradients and point charge gradients
call getQuickEnergyGradients(coord, nxt_charges, xc_coord, &
totEne, gradients, ptchgGrad, ierr)
! Print values obtained from quick library.
#ifdef MPIV
! Dumb way to sequantially print from all cores.
call MPI_BARRIER(MPI_COMM_WORLD,mpierror)
do j=0, mpisize-1
if(j .eq. mpirank) then
call printQuickMPIOutput(natoms, nxt_charges, atomic_numbers, totEne, &
gradients, ptchgGrad, mpirank)
endif
call MPI_BARRIER(MPI_COMM_WORLD,mpierror)
enddo
#else
call printQuickOutput(natoms, nxt_charges, atomic_numbers, totEne, gradients, ptchgGrad)
#endif
! Deallocate memory of point charge stuff.
if ( allocated(xc_coord)) deallocate(xc_coord, stat=ierr)
if ( allocated(ptchgGrad)) deallocate(ptchgGrad, stat=ierr)
enddo
! Finalize QUICK, required only once.
call deleteQuickJob(ierr)
! Deallocate memory.
if ( allocated(atomic_numbers)) deallocate(atomic_numbers, stat=ierr)
if ( allocated(coord)) deallocate(coord, stat=ierr)
if ( allocated(gradients)) deallocate(gradients, stat=ierr)
#ifdef MPIV
call mpi_exit
#endif
end program test_quick_api
Note that in our test program, errors are propagated from QUICK using ierr integer variable. The errors must be properly handled although we have not shown error handling here. Assuming we configured QUICK serial version with a prefix and compiled using intel compiler toolchain,we can compile above source files and link QUICK libraries as follows.
ifort -cpp test_module.f90 example_program.f90 -o example_program -I$installdir/include/serial/
-L$installdir/lib/serial/ -lquick -lblas-quick -lxc -lstdc++
MPI version of the libraries can be linked as follows.
mpiifort -cpp -DMPIV test_module.f90 example_program.f90 -o example_program
-I$installdir/include/mpi/ -L$installdir/lib/mpi/ -lquick-mpi -lblas-quick -lxc -lstdc++
CUDA version of the libraries can be linked as follows.
ifort -cpp test_module.f90 example_program.f90 -o example_program -I$installdir/include/cuda/
-L$installdir/lib/cuda/ -L$CUDA_HOME/lib64 -lcuda -lm -lcudart -lcublas -lcusolver
-lquick-cuda -lxc-cuda -lstdc++
CUDAMPI version of the libraries can be linked as follows.
mpiifort -cpp -DMPIV test_module.f90 example_program.f90 -o example_program
-I$installdir/include/cuda/ -L$installdir/lib/cuda/ -L$CUDA_HOME/lib64 -lcuda -lm -lcudart
-lcublas -lcusolver -lquick-cudampi -lxc-cuda -lstdc++
Running serial or CUDA executable should produce this output. A similar output may be obtained by running MPI or CUDAMPI version with 2 processes.
Adding new basis sets¶
QUICK follows the basis set format established by the Gaussian software. You have to follow this format if you want to construct your own basis set. Established basis sets can be obtained from the basis set exchange web page. In order to add a basis set into QUICK, one should download the basis set in the Gaussian software format and save it in the basis folder. Then, link this basis set to QUICK by updating the basis_link file inside the basis folder. The basis_link file contains a table in the following format.
___________________________________________________________________________
| Keyword | Filename |
|-------------------------------------------------------------------------|
| #STO-3G | STO-3G.BAS |
| #3-21G | 3-21G.BAS |
| #6-31G | 6-31G.BAS |
| #6-31G* | 6-31GS.BAS |
| #6-31G** | 6-31GSS.BAS |
| #6-311G | 6-311G.BAS |
| #6-311G(d,p) | 6-311GDP.BAS |
| #6-311G* | 6-311GS.BAS |
| #6-311G** | 6-311GSS.BAS |
| #cc-pVDZ | CC-PVDZ.BAS |
| #cc-pVTZ | CC-PVTZ.BAS |
|_________________________________________________________________________|
You should update this table by adhering to the rules below.
- Add a keyword for your basis set. This must be followed by a single space and ‘#’ character.
- Keyword size must be less than 32 characters.
- Filename must start at 24th position of the line and must not be longer than 36 characters.
- DO NOT CHANGE THE TABLE/COLUMN WIDTH! VERTICAL BORDERS MUST REMAIN THE SAME.
Note 1: Current version of QUICK (v22.03) ERI engine only support basis functions up to d. Therefore, do not add high angular momentum basis sets and attempt to use f/g functions.
Note 2: ECPs are not supported by QUICK-22.03. Therefore care must be taken not to add elements that require ECPs as this would lead to wrong results.
Adding new test cases into test suite¶
In order to add new test cases into the QUICK test suite, one must follow 3 steps. First, the test input and reference output file should be added into $QUICK_HOME/test and $QUICK_HOME/test/saved directories respectively. Make sure to adhere to following naming convention.
<calculation_type>_<molecule_name>_<QM_method>_<basis_set>.<in|out>
Some example test case names that follow this convention are shown below.
ene_NH4_rhf_631g.in, ene_NH4_rhf_631g.out
grad_CH4_b3lyp_def2svp.in, grad_CH4_b3lyp_def2svp.out
opt_wat_rhf_631g.in, opt_wat_rhf_631g.out
Second, test list files located inside $QUICK_HOME/test should be updated. These are just .txt files that record names of test cases. As of QUICK-22.03, there are 4 testlist files: testlist_short.txt, testlist_short_cuda.txt, testlist_full.txt, testlist_full_cuda.txt. The first and second contain short test lists that would be used for standard testing (i.e. by executing make test or ./runtest commands) of quick/quick.MPI and quick.cuda/quick.cuda.MPI executables. The third and fourth are used for robust testing (i.e. by executing make fulltest or ./runtest - -full commands) which usually happens during CI. They have the following format:
ene_wat2_mp2_631g #MP2 test with s and p basis functions
ene_wat2_mp2_631gss #MP2 test with s, p and d basis functions
grad_psb3_b3lyp_631g #B3LYP gradient test with s and p basis functions
grad_NaCl_b3lyp_def2svp #B3LYP gradient test with s, p and d basis functions
grad_wat_b3lyp_ccpvdz #B3LYP point charge gradient test
opt_wat_rhf_631g #RHF geometry optimization test with s and p basis functions
Third, the runtest script located in $QUICK_HOME/tools should be updated with test information. Specifically, print_test_info() function of the script contains a case statement that sets a string variable value which will be printed during the test runs.
print_test_info(){
.
.
.
case "$t" in
ene_AlH3_rhf_sto3g) testinfo="ALH3: RHF energy test: STO-3G basis set";;
ene_BeH2_rhf_sto3g) testinfo="BeH2: RHF energy test: STO-3G basis set";;
ene_BH3_rhf_sto3g) testinfo="BH3: RHF energy test: STO-3G basis set";;
You should add the new test name as a case and store test information in testinfo variable. Finally, when you add changes using git, make sure to use git add -f command for adding the reference output files. This is due to the fact that files with .out extension are ignored by git according to .gitignore rules.
Maintaining the documentation¶
This section provides some guidence to keep this documentation alive and up to date when the current doc keeper is gone.
The documentation is written in the rst language and you must be familiar with the syntax before starting. A short and sweet rst lesson can be found here.
When you are ready, clone the documentation from GitHub repository.
In the root QUICK-docs directory (from now on $QUICK_DOCS), you should find two directories called docs and resources. Inside the docs directory, a Makefile and source directory should exist. The latter contains all the documentation source and images that would go in. If you have large text files to be included, these should be saved in $QUICK_DOCS/resources inside an appropriate directory and linked properly. Once you have made changes, make sure to set the QUICK version (i.e. set version variable, semantic versioning must be used) in $QUICK_DOCS/docs/source/conf.py and compile the documentation using Make. In order to do so, you must have installed sphinx and python in your system. If the sphix-build executable is not accessible through your path variable, make sure to set the SPHINXBUILD variable in $QUICK_DOCS/docs/Makefile. Then from $QUICK_DOCS/docs folder, execute the following command.
cd $QUICK_DOCS/docs
make html
This should compile the documentation. Once the compilation is done, open up the documentation and check your changes. This may be done as follows.
open $QUICK_DOCS/docs/build/html/index.html
If you are happy with the changes, push/merge the new content into GitHub repository.
The QUICK-docs GitHub repository is linked to readthedocs.org web portal where the documentation is compiled and hosted. Log into merzlab account of the readthedocs.org web portal using appropriate username and password. You should find the following QUICK-docs project page once landed inside the account.
Note that we have different documentation versions in versions in the panel. The latest version is a compilation of the most recent source from the QUICK-docs repository. The other versions correspond to different QUICK release versions.
To create such a version, you must create a GitHub tag that points to a specific commit of the QUICK-docs repository. More details on creating a GitHub tag can be found in the GitHub documentation. Once the tag is created, you can select this tag and create a new documentation version from the versions page of the readthedocs web portal. Note that when creating the GitHub tag, you must name it following semantic versioning. Otherwise, the tag wont appear in the readthedocs web portal. Next, build the documentation by simply hitting the Build Version button of the build page. Once the documentation is built, this will appear online.
You can link the html pages of a particular documentation version anywhere you want. For example, we can link the installation guide of the documentation into the README.md file of QUICK repository.
* [Installation Guide](https://quick-docs.readthedocs.io/en/21.3.0/installation-guide.html#installation-guide)
Note that above we link a page from documentation version 21.3.0 into the QUICK-21.03 README.md file. Similarly, a status badge from a particular version can be included in the README.md file.
Note that you should never link anything from the documentation version named latest. This version will change whenever you make changes to the QUICK-docs repository and thus must be used for testing purposes only.
Last updated by Madu Manathunga on 03/03/2022.